 ClustEval
clustering evaluation framework
ClustEval
clustering evaluation framework
Here you see general information about the clustering method.
Clustering Method 'Hierarchical Clustering' - General
Hierarchical Clustering
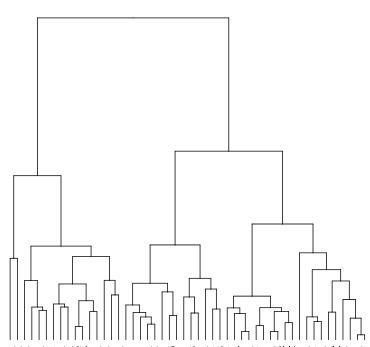 Hierarchical clustering creates a tree from a given set of objects. Each node in the tree represents a cluster. Agglomerative algorithms initially assign all objects to singleton clusters, which are merged based on their similarity (bottom-up). Divisive algorithms start with one big cluster, which is decomposed into smaller clusters along the tree (top-down). A linkage function determines the distance between two clusters. As linkage function, we use the Unweighted Pair Group Method using arithmetic Averages (known as UPGMA or average linking), which is probably the most popular algorithm for hierarchical clustering in computational biology
Hierarchical clustering creates a tree from a given set of objects. Each node in the tree represents a cluster. Agglomerative algorithms initially assign all objects to singleton clusters, which are merged based on their similarity (bottom-up). Divisive algorithms start with one big cluster, which is decomposed into smaller clusters along the tree (top-down). A linkage function determines the distance between two clusters. As linkage function, we use the Unweighted Pair Group Method using arithmetic Averages (known as UPGMA or average linking), which is probably the most popular algorithm for hierarchical clustering in computational biology
- Publication: R Core Team. R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing, Vienna, Austria, 2013. ISBN 3-900051-07-0.
| Information | Value |
|---|---|
| Name | programs/HierarchicalClusteringRProgram.jar |
| Download | Click |
 Hints:
Hints: